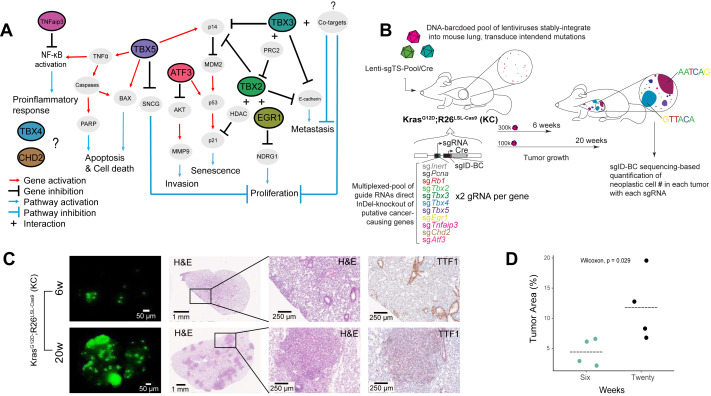
In vivo multiplexed modeling reveals diverse roles of the TBX2 subfamily and Egr1 in Kras-driven lung adenocarcinoma


The TBX2 subfamily of T-box transcription factors (e.g., Tbx2, Tbx3, Tbx4, Tbx5) plays an essential role in lung development. Down-regulation of these genes in human lung adenocarcinoma suggests that these genes may be tumor-suppressive; however, because down-regulation appears to occur primarily via epigenetic change, it remains unclear if these changes causally drive tumor progression or are merely the consequence of upstream events. Herein, we developed the first multiplexed mouse model to study the impact of TBX2 subfamily loss, alongside associated signaling genes (Egr1, Chd2, Tnfaip3a, and Atf3) in Ras-driven lung cancer. Using tumor-barcoding with high-throughput barcode sequencing (TuBa-seq), a high-throughput tumor-barcoding system, we quantified the growth effects of these knockouts during early and late tumorigenesis. Chd2 knockout suppressed both tumor initiation and progression, whereas Tnfaip3 knockout enhanced tumor initiation and overall tumor growth. Tbx2 loss showed stage-specific effects on tumor development. Notably, Egr1 emerged as a strong tumor suppressor and its knockout resulted in approximately a fivefold increase in tumor size at 20 weeks (two-sample t-test, p < 0.05), exceeding the impact observed with Rb1 loss. Transcriptomic analyses of Egr1-deficient tumors suggested immune dysregulation, including heightened inflammation and potential markers of T cell exhaustion in the tumor microenvironment. These findings indicate that Egr1 may play a role in suppressing tumor growth through modulating immune dynamics, offering new insights into the interplay between tumor progression and immune regulation in lung adenocarcinoma.